|
Long-sleep mouse model induced by Sik3 splice mutation C57BL/6N-Sik3<tm1.1Iiis> (RBRC09910)C57BL/6N-Sik3<em2Iiis> (RBRC09911)C57BL/6N-Sik3<em3Iiis> (RBRC09912)Courtesy of Masashi Yanagisawa, M.D., Ph.D.
|
|
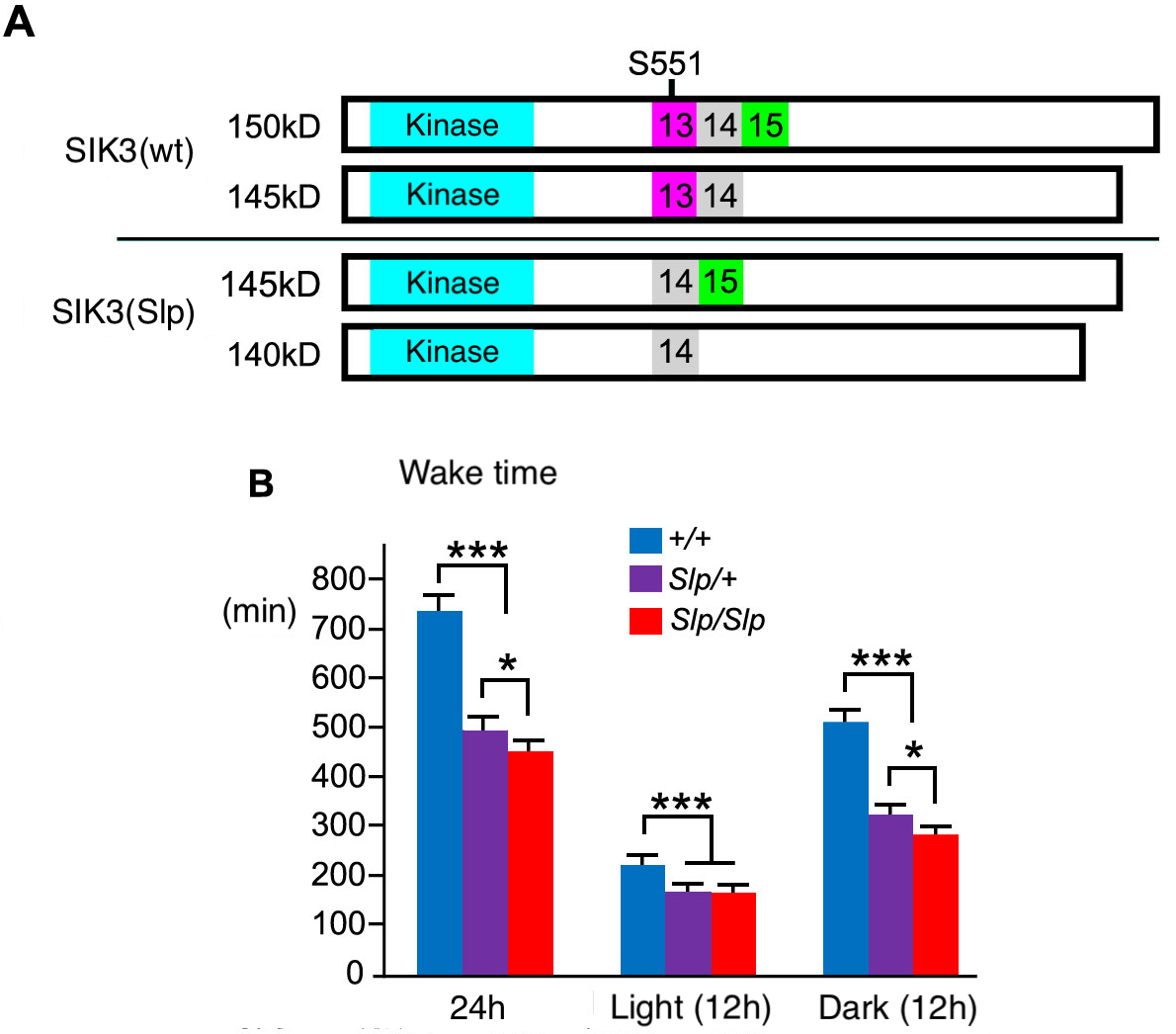
Sleep is an ubiquitously conserved animal behavior. Though sleep is controlled by a strict homeostatic manner, the molecular mechanism is not clear. Depositor (Dr. Yanagisawa) and his colleague found a mutant mouse strain with an unusual long sleep time (Sleepy mice) through a phenotype-driven forward-genetics approach (1). Further analysis for Sleepy mice identified a novel sleep control factor Sik3. SIK3 belongs to AMP-activated protein kinase-related kinase family and broadly express in neurons. Sleepy mice have a splicing mutation in Sik3 protein kinase gene (Sik3(SLP)). Knock-in mice with Sik3(SLP) allele show the similar phenotype to original Sleepy mice. The relationship between Sik3 and sleep is a hot topic now. For examples, Sik3(SLP) mice analysis clarified the existence of sleep-need-index phosphoproteins (SNIPPs) (2). Moreover, it was reported that a single amino acid S551 of SIK3 is important for sleep-wake regulation (3). Not only mutant Sik3(SLP) mice (RBRC09910), but also wild-type Sik3 Flag-HA Tag KI mice (RBRC09911) and Sik3(SLP) Flag-HA Tag KI mice (RBRC09912) are available from RIKEN BioResource Research Center. These Flag-HA Tag KI strain are especially useful for protein analysis. |
| Depositor | : | Masashi Yanagisawa, M.D., Ph.D. International Institute for Integrative Sleep Medicine, University of Tsukuba |
|
| Strain name | : | C57BL/6N-Sik3<tm1.1Iiis> | |
| RBRC No. | : | RBRC09910 | |
| Strain name | : | C57BL/6N-Sik3<em2Iiis> | |
| RBRC No. | : | RBRC09911 | |
| Strain name | : | C57BL/6N-Sik3<em3Iiis> | |
| RBRC No. | : | RBRC09912 | |
| References | : | (1) | Funato H, Miyoshi C, Fujiyama T, Kanda T, Sato M, Wang Z, Ma J, Nakane S, Tomita J, Ikkyu A, Kakizaki M, Hotta-Hirashima N, Kanno S, Komiya H, Asano F, Honda T, Kim SJ, Harano K, Muramoto H, Yonezawa T, Mizuno S, Miyazaki S, Connor L, Kumar V, Miura I, Suzuki T, Watanabe A, Abe M, Sugiyama F, Takahashi S, Sakimura K, Hayashi Y, Liu Q, Kume K, Wakana S, Takahashi JS, Yanagisawa M. Forward-genetics analysis of sleep in randomly mutagenized mice. Nature.; 539(7629):378-383, 2016. |
| (2) | Wang Z, Ma J, Miyoshi C, Li Y, Sato M, Ogawa Y, Lou T, Ma C, Gao X, Lee C, Fujiyama T, Yang X, Zhou S, Hotta-Hirashima N, Klewe-Nebenius D, Ikkyu A, Kakizaki M, Kanno S, Cao L, Takahashi S, Peng J, Yu Y, Funato H, Yanagisawa M, Liu Q. Quantitative phosphoproteomic analysis of the molecular substrates of sleep need. Nature.; 558(7710):435-439, 2018. | ||
| (3) | Honda T, Fujiyama T, Miyoshi C, Ikkyu A, Hotta-Hirashima N, Kanno S, Mizuno S, Sugiyama F, Takahashi S, Funato H, Yanagisawa M. A single phosphorylation site of SIK3 regulates daily sleep amounts and sleep need in mice. Proc Natl Acad Sci U S A.; 115(41):10458-10463, 2018. | ||
| March 2019 Contact: Saori Mizuno, Ph.D. Experimental Animal Division, RIKEN BioResource Research Center All materials contained on this site may not be reproduced, distributed, displayed, published or broadcast without the prior permission of the owner of that content. |